OENOLOGICAL POTENTIAL OF SACCHAROMYCES CEREVISIAE AND KLUYVEROMYCES MARXIANUS STRAINS FOR USE IN MIXED FERMENTATIONS
Keywords:
Kluyveromyces marxianus, Saccharomyces cerevisiae, wine, mixed culture, oenological characteristics.Abstract
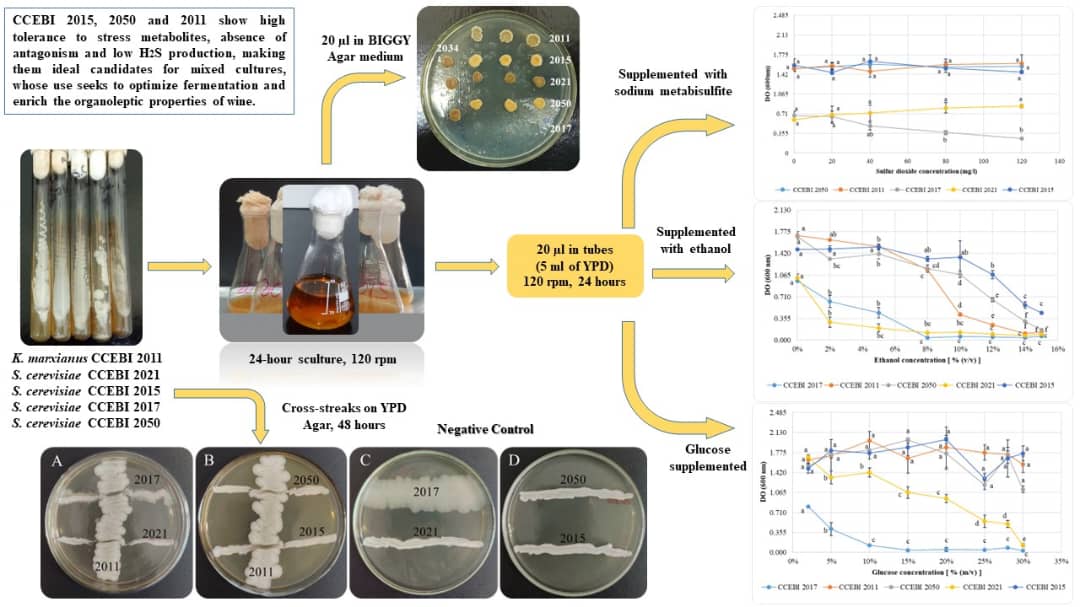
Wine fermentation is a complex process involving several yeast species. Saccharomyces cerevisiae species is particularly effective due to its fermentative efficiency, while non-Saccharomyces yeasts contribute to the aromatic complexity of wines. This study aimed to characterize five strains (four S. cerevisiae and one Kluyveromyces marxianus), from different isolation sources, in terms of characteristics of oenological interest. The results showed that the CCEBI 2015 and CCEBI 2050 strains from S. cerevisiae species, and K. marxianus CCEBI 2011 tolerate high concentrations of glucose (30 % w/v), ethanol (10 % v/v) and SO₂ (120 mg/l) and have low or no H₂S production. No antagonism was observed between K. marxianus and S. cerevisiae strains, suggesting their compatibility for mixed fermentations. These results indicate that these strains are promising candidates for use in mixed starter cultures for winemaking.
References
FAZIO, N. A. et al. “Inside current winemaking challenges: Exploiting the potential of conventional and unconventional yeasts”. Microorganisms. 2023, 11(5), 1338. https://doi.org/10.3390/microorganisms11051338
COMITINI, F. et al. “Yeast interactions and molecular mechanisms in wine fermentation: a comprehensive review”. Int. J. Mol. Sci. 2021, 22(14), 7754. https://doi.org/10.3390/ijms22147754
ROMANO, P.; MAURIZIO, C.; GRAHAM, H. F. (eds.) Yeasts in the Production of Wine. New York, NY, USA: Springer. 2019, pp. 515. ISBN: 978-1-4939-9782-4. https://doi.org/10.1007/978-1-4939-9782-4
GERŐCS, A. et al. “Characterization of Saccharomyces strains isolated from “Kéknyelű” grape must and their potential for wine production”. Fermentation. 2022, 8(8), 416. https://doi.org/10.3390/fermentation8080416
BARONE, E. et al. “Use of Kluyveromyces marxianus to increase free monoterpenes and aliphatic esters in white wines”. Fermentation. 2021, 7(2), 79. https://doi.org/10.3390/fermentation7020079
VALLEJO-VIDAL, J. A. et al. “A novel Kluyveromyces marxianus strain with an inducible flocculation phenotype”. AMB Express. 2012, 2(38). DOI: https://doi.org/10.1186/2191-0855-2-38
SERRAT-DÍAZ, M. et al. “Influencia de las condiciones de cultivo sobre el crecimiento y contenido de pared celular en una cepa floculante de Kluyveromyces marxianus”. Revista Cubana de Química. 2017, 29(1), 89-102. http://scielo.sld.cu/pdf/ind/v29n1/ind07117.pdf
BILAL, M. et al. “Bioprospecting Kluyveromyces marxianus as a robust host for industrial biotechnology”. Frontiers in Bioengineering and Biotechnology. 2022, 10, 851768. DOI: https://doi.org/10.3389/fbioe.2022.851768
ERASMUS, B.; DIVOL, B. “Exploring the phenotypic diversity of oenological traits in Kluyveromyces marxianus strains”. FEMS Yeast Research. 2022, 22(1), foac009. https://doi.org/10.1093/femsyr/foac009
RUIZ, J. et al. “Occurrence and enological properties of two new non-conventional yeasts (Nakazawaea ishiwadae and Lodderomyces elongisporus) in wine fermentations”. International Journal of Food Microbiology. 2019, 305, 108255. DOI:
https://doi.org/10.1016/J.IJFOODMICRO.2019.108255
ESCRIBANO-VIANA, R. et al. “Selection process of a mixed inoculum of non-Saccharomyces yeasts isolated in the DO Ca. Rioja”. Fermentation. 2021, 7(3), 148. https://doi.org/10.3390/fermentation7030148
CORDENTE, A. G. et al. “Isolation of sulfite reductase variants of a commercial wine yeast with significantly reduced hydrogen sulfide production”. FEMS Yeast Research. 2009, 9(3), 446-459. https://doi.org/10.1111/j.1567-1364.2009.00489.x
LERTCANAWANICHAKUL, M.; SAWANGNOP, S. A “Comparison of two methods used for measuring the antagonistic activity of Bacillus species”. Walailak Journal of Science and Technology. 2008, 5(2):161-167. https://wjst.wu.ac.th/index.php/wjst/article/view/86
D'AMATO, D. et al. “Effects of temperature, ammonium and glucose concentrations on yeast growth in a model wine system”. Int. J. Food Sci. Tech. 2006, 41, 1152-1157. DOI: https://doi.org/10.1111/j.1365-2621.2005.01128.x
LAPPE-OLIVERAS, P. et al. “Genotypic and phenotypic diversity of Kluyveromyces marxianus isolates obtained from the elaboration process of two traditional mexican alcoholic beverages derived from Agave: Pulque and Henequen (Agave fourcroydes) Mezcal”. Journal of Fungi. 2023, 9(8), 795. https://doi.org/10.3390/jof9080795
FERREIRA, J.; TOIT, M. D.; TOIT, W. D. “The effects of copper and high sugar concentrations on growth, fermentation efficiency and volatile acidity production of different commercial wine yeast strains”. Australian Journal of Grape and Wine Research. 2006, 12(1), 50-56. Doi: https://doi.org/10.1111/j.1755-0238.2006.tb00043.x
SIPICZKI, M. “Yeast two- and three-species hybrids and high-sugar fermentation”. Microbial Biotechnology. 2019, 12(6), 1101-1108. https://doi.org/10.1111/1751-7915.13390
YOSHIDA, M. et al. “Wine yeast cells acquire resistance to severe ethanol stress and suppress insoluble protein accumulation during alcoholic fermentation”. Microbiology Spectrum. 2022, 10(5), e00901-22. https://doi.org/10.1128/spectrum.00901-22
MA, M.; LIU, Z. L. “Mechanisms of ethanol tolerance in Saccharomyces cerevisiae”. Applied Microbiology and Biotechnology. 2010, 87, 829-845. https://doi.org/10.1007/s00253-010-2594-3
MARULLO, P. et al. “Natural allelic variations of Saccharomyces cerevisiae impact stuck fermentation due to the combined effect of ethanol and temperature; a QTL-mapping study”. BMC genomics. 2019, 20, 1-17. https://doi.org/10.1186/s12864-019-5959-8
LAIRÓN-PERIS, M. et al. “Lipid composition analysis reveals mechanisms of ethanol tolerance in the model yeast Saccharomyces cerevisiae”. Applied and Environmental Microbiology. 2021, 87(12), e00440-21. https://doi.org/10.1128/AEM.00440-21
FENG, L. et al. “Selection of indigenous Saccharomyces cerevisiae strains for winemaking in Northwest China”. American Journal of Enology and Viticulture. 2019, 70(2), 115-126. DOI: https://doi.org/10.5344/ajev.2018.18035
CAMACHO-POZO, M. I. et al. “Evaluation of two conservation methods for Kluyveromyces marxianus CCEBI 2011 at the CEBI Culture Collection”. Revista de la Sociedad Venezolana de Microbiología. 2014, 34, 91-96. https://ve.scielo.org/pdf/rsvm/v34n2/art09.pdf
KARIM, A.; GERLIANI, N.; AÏDER, M. “Kluyveromyces marxianus: An emerging yeast cell factory for applications in food and biotechnology”. International Journal of Food Microbiology. 2020, 333, 108818. https://doi.org/10.1016/j.ijfoodmicro.2020.108818
ZARA, G.; NARDI, T. “Yeast metabolism and its exploitation in emerging winemaking trends: From sulfite tolerance to sulfite reduction”. Fermentation. 2021, 7, 57. https://doi.org/10.3390/fermentation7020057
MOLINA-ESPEJA, P. “Next generation winemakers: Genetic engineering in Saccharomyces cerevisiae for trendy challenges”. Bioengineering. 2020, 7(4), 128. https://doi.org/10.3390/bioengineering7040128
DIVOL, B.; DU TOIT, M.; DUCKITT, E. “Surviving in the presence of sulphur dioxide: Strategies developed by wine yeasts”. Appl. Microbiol. Biotechnol. 2012, 95, 601-613. https://doi.org/10.1007/s00253-012-4186-x
DE GUIDI, I. et al. “Development of a new assay for measuring H2S production during alcoholic fermentation: Application to the evaluation of the main factors impacting H2S production by three Saccharomyces cerevisiae wine strains”. Fermentation. 2021, 7(4), 213. https://doi.org/10.3390/fermentation7040213
LI, Y. et al. “Saccharomyces cerevisiae isolates with extreme hydrogen sulfide production showed different oxidative stress resistances responses during wine fermentation by RNA sequencing analysis”. Food Microbiology. 2019, 79, 147-155. https://doi.org/10.1016/j.fm.2018.10.021
ALTURKI, S. N. et al. “Killer phenomenon in yeast: An overview”. Journal of American Science. 2019, 15(4). Doi:
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Rolando Carrazana-Isaac, Manuel de J. Serrat-Díaz

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
This journal provides immediate open access to its content, based on the principle that offering the public free access to research helps a greater global exchange of knowledge. Each author is responsible for the content of each of their articles.






















